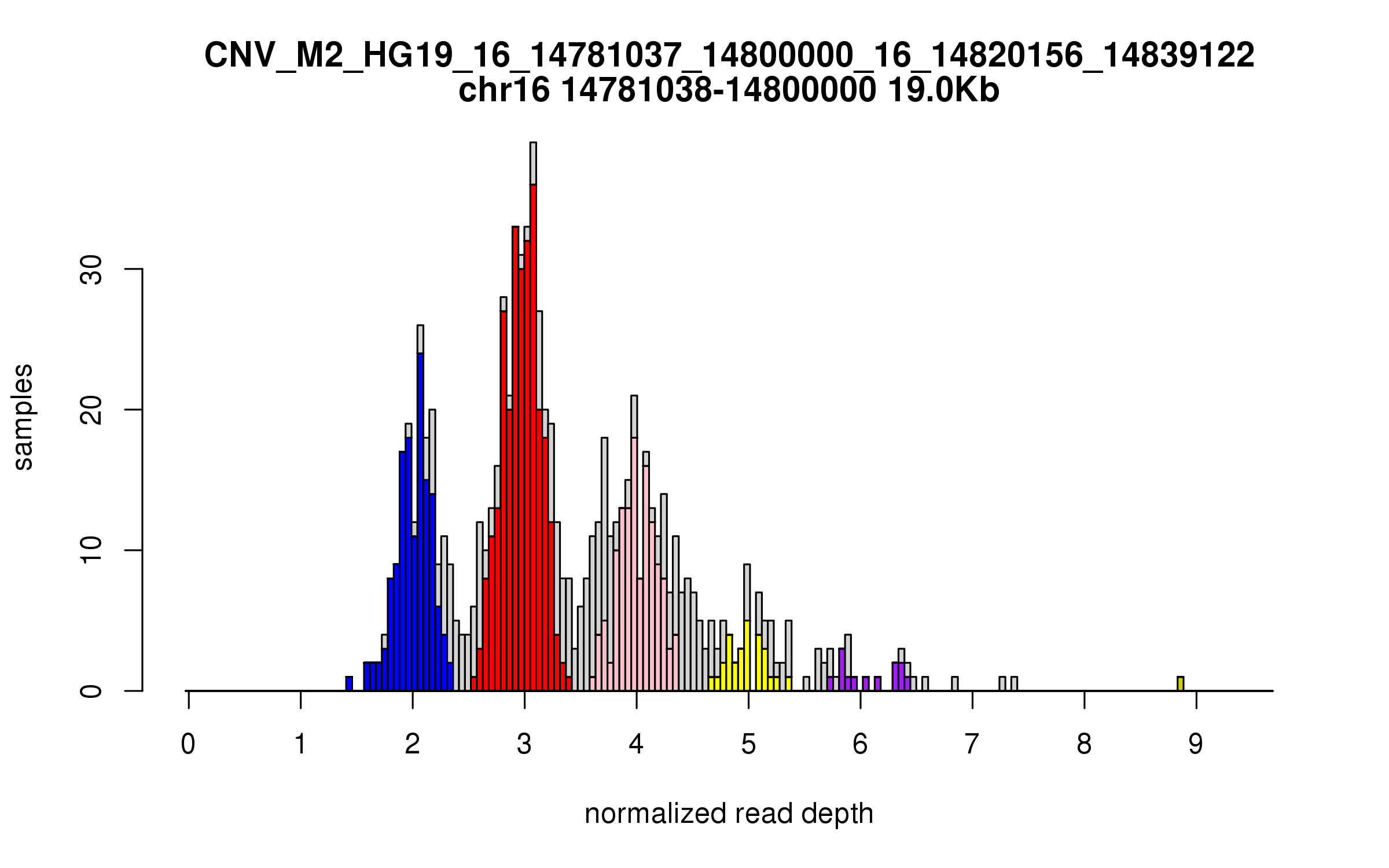
Variant CNV_M2_HG19_16_14781037_14800000_16_14820156_14839122
Location: chr16:14781037-14800000 (18Kb), chr16:14820156-14839122 (18Kb), distance: 20Kb
Site classification: Multi-allelic
Variant DCN frequency: 0.530
Genes Overlapped
Site classification: Multi-allelic
Variant DCN frequency: 0.530
Genes Overlapped
| PLA2G10 | CDS | group 10 secretory phospholipase A2 precursor |
| RP11-719K4.1 | EXON | NA |
1000 Genomes Phase 1
Diploid copy number distribution
POP CN0 CN1 CN2 CN3 CN4 CN5 CN6 CN7 CN8 CN9
ASW 0 0 4 12 19 8 1 0 0 1
CEU 0 0 24 34 7 1 0 0 0 0
CHB 0 0 11 31 29 7 0 0 0 0
CHS 0 0 8 49 27 6 1 0 0 0
CLM 0 0 12 21 14 2 0 0 0 0
FIN 0 0 31 26 4 1 0 0 0 0
GBR 0 0 15 28 11 0 0 0 0 0
IBS 0 0 2 3 1 0 0 0 0 0
JPT 0 0 6 32 23 5 3 0 0 0
LWK 0 0 0 20 33 7 8 1 0 0
MXL 0 0 10 23 16 4 1 0 0 0
PUR 0 0 16 27 9 0 0 0 0 0
TSI 0 0 31 38 13 1 0 0 0 0
YRI 0 0 2 10 24 20 14 1 0 0
AFR 0 0 2 30 57 27 22 2 0 0
AFR+ 0 0 6 42 76 35 23 2 0 1
AMR 0 0 38 71 39 6 1 0 0 0
ASN 0 0 25 112 79 18 4 0 0 0
EUR 0 0 103 129 36 3 0 0 0 0
ALL 0 0 172 354 230 62 28 2 0 1
Diploid copy number distribution (95% confident)
POP CN0 CN1 CN2 CN3 CN4 CN5 CN6 CN7 CN8 CN9
ASW 0 0 1 9 11 1 1 0 0 1
CEU 0 0 23 27 4 0 0 0 0 0
CHB 0 0 7 23 18 4 0 0 0 0
CHS 0 0 6 29 5 2 1 0 0 0
CLM 0 0 11 17 5 1 0 0 0 0
FIN 0 0 25 20 1 0 0 0 0 0
GBR 0 0 9 20 8 0 0 0 0 0
IBS 0 0 0 2 0 0 0 0 0 0
JPT 0 0 5 30 18 3 1 0 0 0
LWK 0 0 0 10 14 2 1 0 0 0
MXL 0 0 6 16 9 1 0 0 0 0
PUR 0 0 13 21 2 0 0 0 0 0
TSI 0 0 29 37 10 0 0 0 0 0
YRI 0 0 2 9 20 14 9 0 0 0
AFR 0 0 2 19 34 16 10 0 0 0
AFR+ 0 0 3 28 45 17 11 0 0 1
AMR 0 0 30 54 16 2 0 0 0 0
ASN 0 0 18 82 41 9 2 0 0 0
EUR 0 0 86 106 23 0 0 0 0 0
ALL 0 0 137 270 125 28 13 0 0 1
Imputation spider plots: ASW CEU CHB CHS CLM FIN GBR IBS JPT LWK MXL PUR TSI YRI